It’s been a few months since our November Update and there is a lot to share: we have several new preprints on our wastewater work, scaled up our nasal swab collection dramatically, we’re moving into a larger lab space, and we’re now called “SecureBio Detection”. As always, if you have questions or see opportunities to collaborate, please let us know; we’re eager to work with others thinking along similar lines.
Wastewater Sequencing
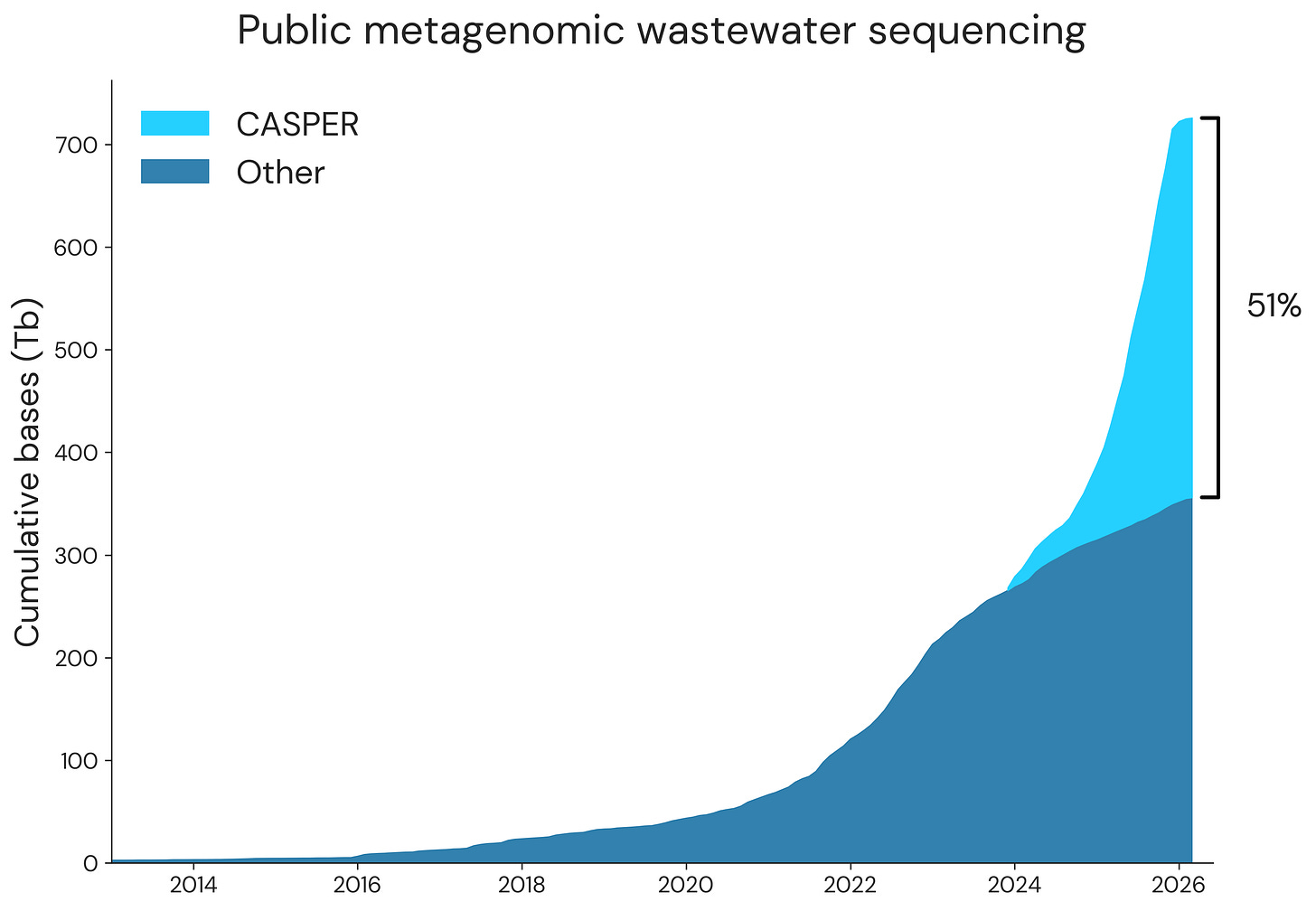
We’ve continued to collaborate with our partners across CASPER to sequence and analyze municipal wastewater, and recently shared a preprint describing our progress so far. We also hit a significant milestone, where the majority of metagenomic wastewater sequencing data on SRA is from CASPER:
This data, all 1.3 trillion sequencing reads, is available at PRJNA1247874, and we’re excited to see what people will do with this rich dataset!
Our paper describing our collaboration with Jason Rothman, Katrine Whiteson, and the Southern California Coastal Water Research Project to sequence wastewater from Los Angeles has now been published in Nature Scientific Data. This peer-reviewed publication is an update to research we shared as a preprint in March 2025.
Rachel Poretsky wrote a case study that she submitted to NEJM Evidence, describing the first CASPER detection of wild-type measles. This represented the identification of a single case out of a sewershed of over 1M people. While this is promising, to fully understand sensitivity to measles we would need to assess how many of our monitored sewersheds had cases of measles that we did not detect. We’ve since detected measles on a total of 21 occasions.
While we’re all working to get the end-to-end time down in the typical case, CASPER recently hit a new “personal best” of seven days: we enabled Marc Johnson to notify a local public health partner on 2026-03-16 about a detection of measles in a sample collected on 2026-03-09. We see a path to get the end-to-end time down to five days, and with aggressive investment eventually three, and we’re actively fundraising for this work.
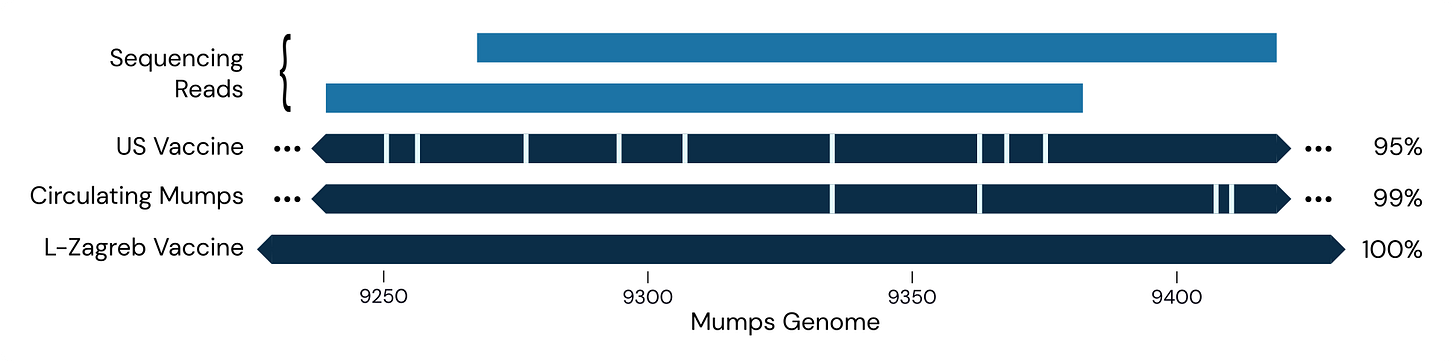
In November we detected mumps in one of the CASPER sites downstream from an international airport, but it had several genomic mismatches relative to circulating strains and was even more distant from the Jeryl Lynn vaccine strains used in the US. It was, however, an exact match to the L-Zagreb vaccine strain, which is used outside the US:
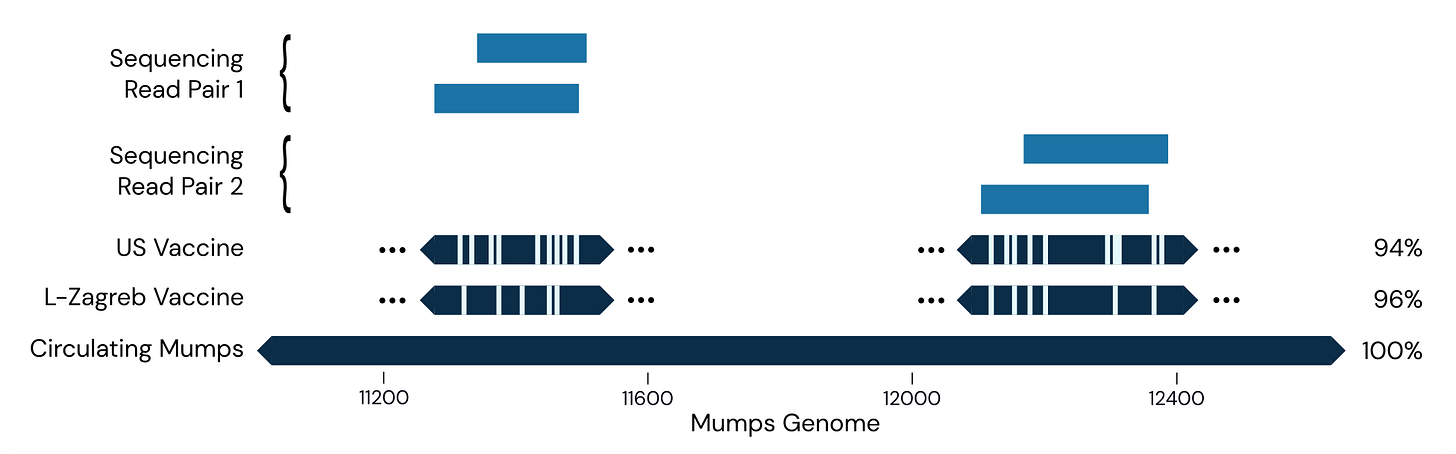
This suggests we picked up someone who was vaccinated shortly before flying here. More recently, however, we had our first detection of wild-type mumps, a 100% match for mumps genotype C:
This level of precision, with strain-level identification, illustrates just how rich sequencing data is.
Pooled Individual Sequencing (Zephyr)
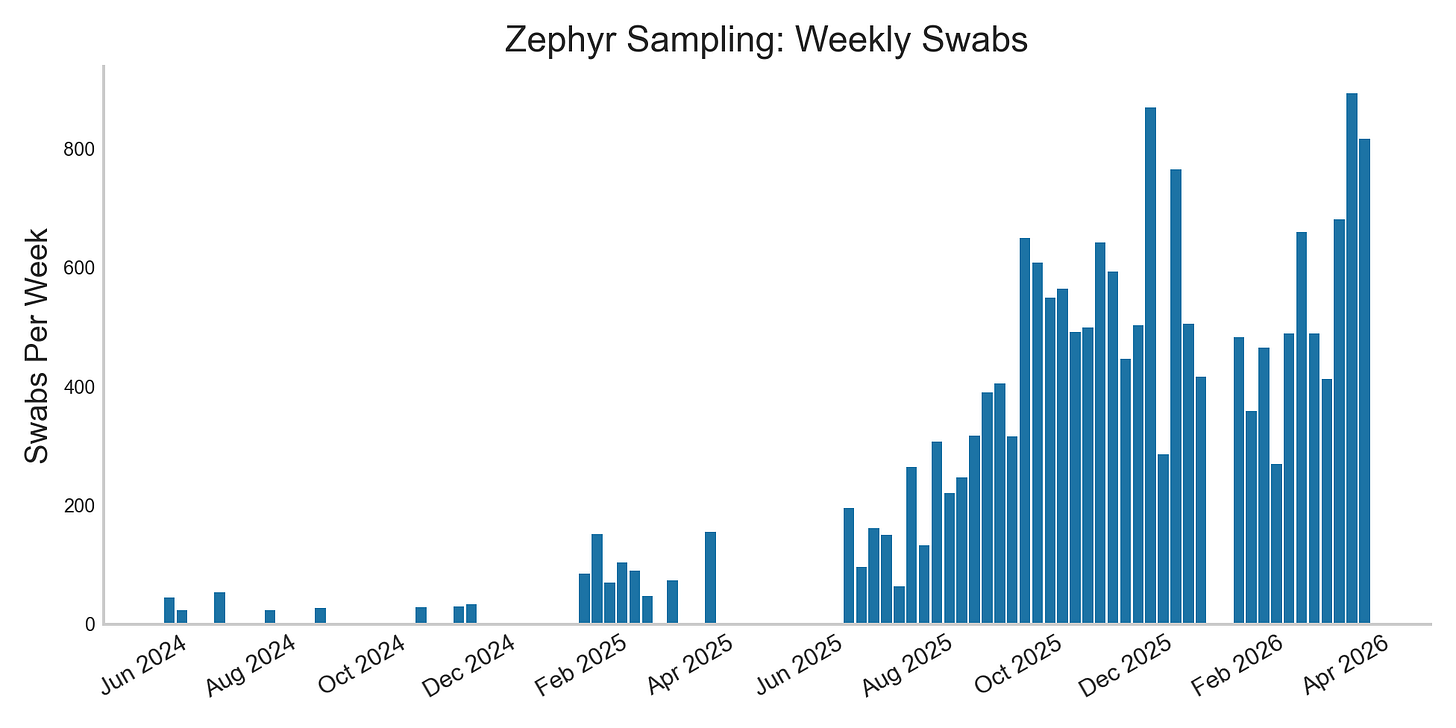
We’ve continued Zephyr, our Boston-based swab sampling program, with dedicated field samplers going out most days. We shared a blog post describing our work, and were featured in the Boston Globe.
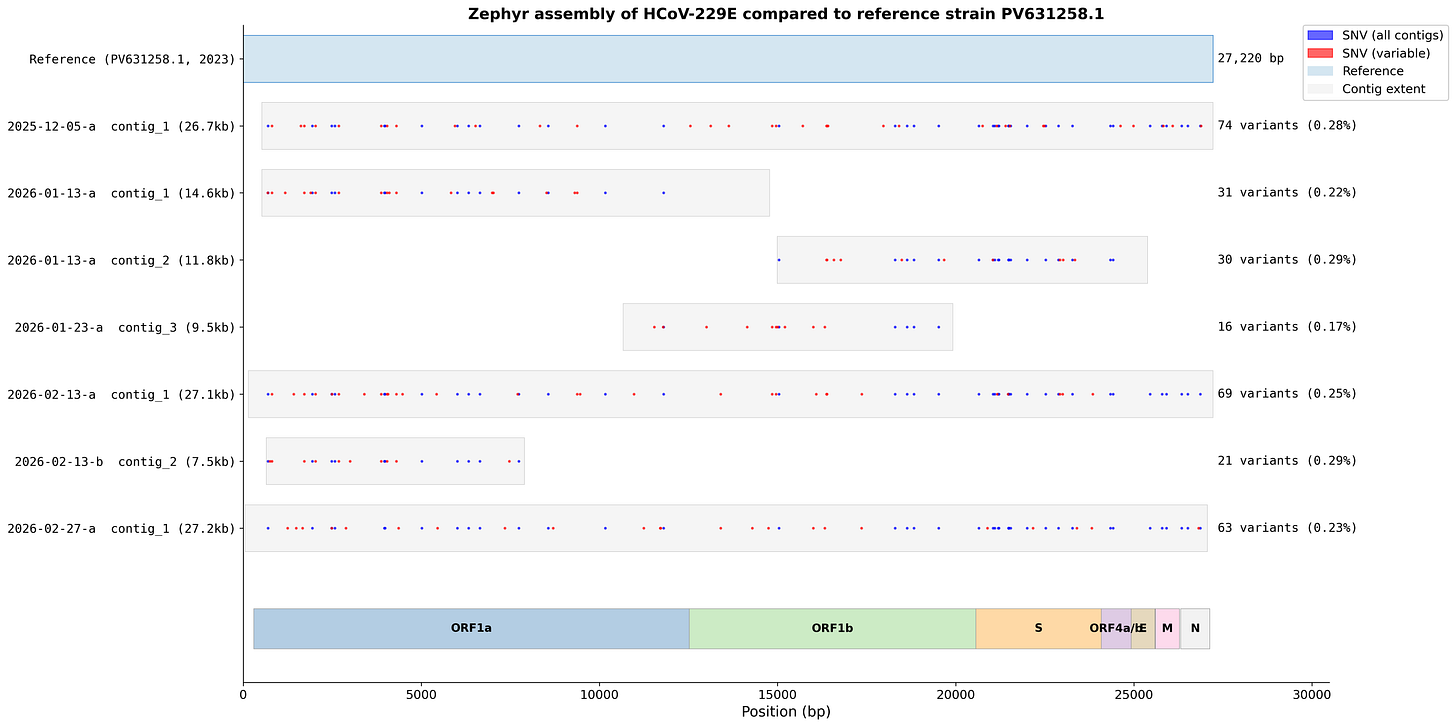
We’ve now collected over 16,000 swabs, 75% of which during this respiratory season. In these we’ve detected a wide variety of respiratory viruses, including four separate strains of non SARS-CoV-2 coronaviruses, and over a dozen rhinovirus serotypes. For many of these, using Oxford Nanopore Sequencing has allowed us to recover near-complete genomes. These genomes identify the specific cold and flu strains spreading in Boston, and how they differ from other regions. Here’s an example showing how we’re seeing a consistent local variant of the coronavirus 229E, across 6 sequencing runs:
We also ran an initial experiment on bacterial sequencing from nasal swabs. While it was promising, we’ve had to set this work aside for now due to capacity constraints. We continue to think that sequencing pooled nasal swabs could be an excellent way to detect outbreaks of respiratory bacteria.
Laboratory Work
When we spun out of MIT we moved into Tufts Launchpad Biolabs, in Chinatown. It has been a great home for us, but as our team has continued to grow we’ve decided to move out into our own space. Starting April 1st we will have 7,500 sqft of lab space in Kendall Sq, a short walk from our One Broadway office, which positions us well for further growth.
We’re exploring the extent to which we can automate portions of our sample processing pipeline, which will free up staff for other work, reduce end-to-end time, and increase consistency. At a high level, our approach is to identify the portions of our work that are most amenable to automation and deploy it strategically, as opposed to attempting to automate the entire lab process end-to-end. Freeing up our scientists from routine production workflows by incorporating laboratory automation will also enable us to focus more on R&D work to improve the sensitivity of our current RNA viral detection methods and to bolster pathogen detection in other domains.
We still see PCR biosurveillance as a powerful tool both for public health and as a key component of an emergency response strategy. There are cases, however, where PCR-based detection can fail to detect its intended targets, or fail to provide the level of granularity needed to support rapid decision-making. We describe these in a recent blog post. One area where PCR is unmatched is in its ability to rapidly scale detection for a specific target, and we’re discussing approaches to reduce the time from identifying a novel threat until deploying a validated PCR test
Sampling Strategies
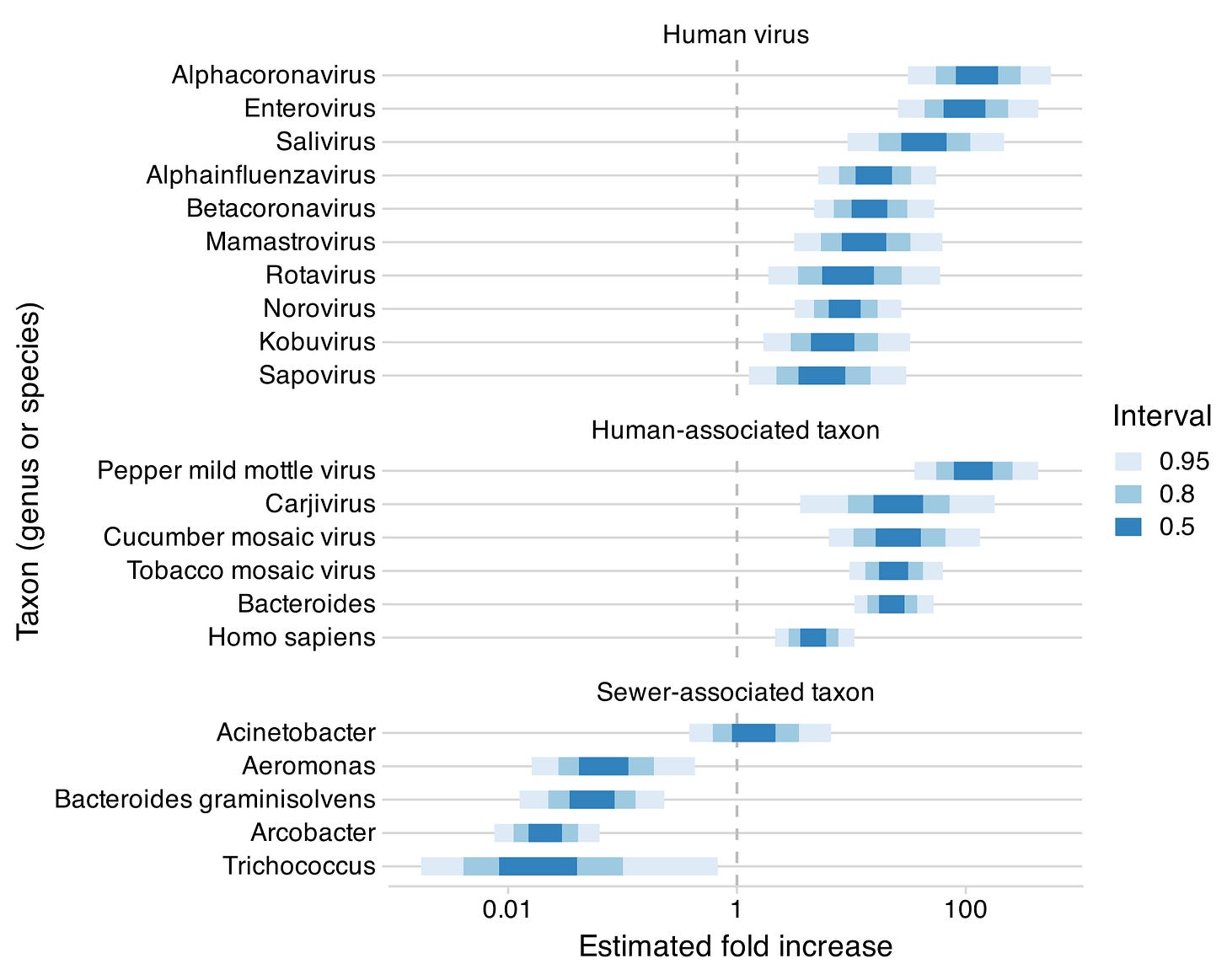
We have a preprint (blog post) on our work in collaboration with CDC and Ginkgo, comparing airplane lavatory vs municipal wastewater. While airplane wastewater is already exciting from an epidemiological perspective, since it reflects travelers, it’s also very promising compositionally: human viruses and other human-associated species were a much larger fraction of the sequencing reads. This is a preliminary result, but we’re working to get access to additional airplane wastewater for further testing.
We also have a preprint on blood supply monitoring, following up on our blog posts (part 1, part 2, part 3) from last year. While we think wastewater and nasal swabs are the highest priority sample types to explore, it’s likely that a fully comprehensive system will need to go beyond these two, and blood is high on the list.
Analysis of Sequencing Data
We collaborated with the AI side of SecureBio to automate initial human review of flags. Our existing pipeline flags a small fraction of reads for human review, but as the number of reads we’re processing has increased that small fraction has threatened to overwhelm our human reviewers. Additionally, human review requires expert judgement, which trades off against time improving our detection capabilities. We now supply frontier models with flagged sequencing reads, bioinformatic annotation, the ability to query BLAST, and a clear rubric, and they handle clear-cut cases quickly and efficiently. This has reduced the human review load by about 80%.
We improved our open source sample characterization and viral identification pipeline, mgs-workflow, in several areas: over the last few months we built out comprehensive continuous integration, a dramatically streamlined release process, several major memory improvements, and schemas for all output files, with documentation and automated enforcement.
We’ve also made large strides in automating the earlier stages of this pipeline, with the goal that the entire flow should happen without any human intervention. This has included pre-populating metadata and automating data import, mgs-workflow’s RUN and DOWNSTREAM stages, and automated partner report generation.
The second half of our chimera detection implementation had some algorithmic inefficiencies, and as our data volume has increased its runtime had ballooned from seconds to hours. Over the past few months we have reduced runtime to a few minutes.
Organizational Updates
We’ve rebranded from the Nucleic Acid Observatory to SecureBio Detection (announcement). Our group first started as a project within MIT’s Sculpting Evolution lab and then became a collaboration between the lab and SecureBio. During that period, it was useful to have independent branding, but we finished spinning out from MIT over a year ago to be fully under SecureBio. Updating our name makes it easier for people to understand what we do and how the pieces of SecureBio fit together.
Kelly Chafin spoke about biodefense and the importance of pathogen-agnostic biosurveillance at both the “Biosecurity 2026: New Challenges, New Opportunities?” workshop organized by the Center for Global Security Research at Lawrence Livermore National Laboratory and the Pandemic Center at Brown University’s webinar on The State of U.S. Biodefense. She also participated in an AIxBio mitigations workshop co-hosted by RAND and Helena.
Siham Elhamoumi, as Head of Partnerships, has been engaging potential partners and stakeholders across the public health and biosecurity community, and we plan to share more about our work here once it’s farther along. If you see opportunities to collaborate with SecureBio, please reach out: siham@securebio.org.
Over the winter we hired several people across the computational and laboratory teams:
Alessandro Zulli, Research Scientist. Alessandro comes to us from the Boehm lab at Stanford University where he was a senior research engineer for WastewaterSCAN, developing wet lab methods and bioinformatics pipelines. At Stanford, he developed novel assays for the detection of emerging pathogens, leading interdisciplinary teams for their nationwide deployment and coordinating responses with local public health. He also worked to expand wastewater based epidemiology internationally, leading a project focused on HIV and TB in South Africa. Most recently, he spearheaded efforts to use metagenomic sequencing of wastewater as a complementary signal to targeted molecular methods. Alessandro earned his PhD at Yale University under Professor Jordan Peccia, where he led development on Connecticut’s wastewater SARS-CoV-2 initiative. He joins us to lead our Zephyr nasal swab sampling program.
Jo Faraguna, Bioinformatics Engineer. Jo comes to us from Ginkgo Bioworks where she was a computational protein engineer, building pipelines and applying them to engineer enzymes and protein binders for industrial applications. Previously, Jo also worked at Ginkgo Bioworks as a yeast metabolic strain engineer and before that received her B.S. in Biological Engineering from MIT. She joins us to design, maintain, and operate our bioinformatic analysis pipelines for high-throughput analysis of metagenomic biosurveillance data. We’re also hoping to draw on her experience with protein engineering as we expand our computational detection methods into amino acid- and protein structure-informed spaces.
Jake Lloyd, Research Associate. Jake is now joining us full time, after almost a year on Zephyr as a Field Sampler. He previously interned in Flagship’s Pioneering Medicine, where he served on the in vivo research team in running and developing protein assays on one of their drug development teams, utilizing eRNA therapeutics to treat chronic diseases. Before that he interned at Solarea Bio, assisting in stability studies for proprietary microbiological products in late-stage development for commercial release. He has recently finished his Masters of Science in Biotechnology at Northeastern University, and will be joining our lab team. Outside of work, Jake is an avid writer and enjoys spending time at the gym.
Jared Gurzenda, Senior Research Associate. Jared joins us from Inari Agriculture, where he led and executed molecular biology workflows, including DNA/RNA extraction, gene expression analysis, next-generation sequencing using Illumina and Oxford Nanopore platforms, and whole plasmid sequencing. His work focused on advancing gene-edited seed development. At Inari, Jared played a key role in scaling operations and integrating automation through liquid handling and worklisting systems, improving sample processing efficiency while maintaining high data quality and reducing FTE requirements. Outside the lab, Jared enjoys exploring nature, identifying native plants and fungi, and trail running with his dog.
Matt Benczkowski, Associate Scientist. Matt joins SecureBio from Tessera Therapeutics, where he developed and optimized genomic and molecular assays supporting a range of therapeutic programs. He brings several years of hands-on experience in sequencing across multiple platforms, as well as expertise in high-throughput dPCR workflows. Matt holds a Master of Public Health in Epidemiology of Microbial Disease from the Yale School of Public Health where he contributed to NIH-supported genomic tracing efforts during the COVID-19 pandemic by applying sequencing and transmission dynamic modeling to better understand disease spread. Previously at the University of Pittsburgh, he worked extensively on bacteriophage discovery and genetic engineering.
We are also close to wrapping up our Metagenomics Scientist hiring round, and we appreciate all the great referrals people have sent us over the past few months!